What'sNEW Archives, January-February 2006
 28 February 2006 28 February 2006
Can Viruses Make Us Human? This is the title of a paper presented to the American Philosophical Society by Luis P. Villarreal, the Director of the Center for Virus Research at the University of California, Irvine. Even we, already convinced of the importance of viruses in evolution, are amazed by it. This essay will develop and present the argument that such stable persisting viruses represent a major creative force in the evolution of the host, driving the host to acquire new, and accumulate ever more complex, molecular identities. Based on this premise, this essay will examine the possible role of viruses in the evolution of complexity.... The traits that viral genes appear to have provided to eukaryotes include

- The eukaryotic nucleus,
- Flowering plants,
- The adaptive immune system in animals,
- Live birth (viviparous) placental mammals,
- The multi-membrane bound separation of transcription from replication,
- Pore structures that actively transport RNA into the cytoplasm,
- Chromatin proteins,
- Linear chromosomes with telomere ends,
- DNA-dependent RNA polymerase,
- The enzymes that modify mRNA in a eukaryotic-specific way,
- The complex role of tubulin in the separation of eukaryotic daughter chromosomes and
- The dissolution and reformation of the nuclear membrane.
Some of the supplied proteins have functions that are clear only in the eukaryotic context, and unexplained in the virus or even its usual host, which may be a bacterium. In standard darwinian theory this surprise is mitigated if viruses stole the genes from eukaryotes in the distant past. But Villarreal thinks the genes went the opposite way: viruses supplied them to eukaryotes. Frequently, the viral version of a gene represents the simplest example of any related protein in a functional protein family. ...A properly conducted phylogenetic analysis will usually show that the viral version is basal to the version found in the host. The viral version appears to be older, often simpler.
If viruses are the earliest source for eukaryotic genes, the standard neo-darwinian account for those genes does not work. But eukaryotic genes supplied by viruses are a basic prediction of cosmic ancestry.
 Luis P. Villarreal, "Can Viruses Make Us Human?", p 296-323 v 148 n 3, Proc Am Phil Soc, Sep 2004
[Research Gate]. Luis P. Villarreal, "Can Viruses Make Us Human?", p 296-323 v 148 n 3, Proc Am Phil Soc, Sep 2004
[Research Gate].
 Greg Bear, "When Genes Go Walkabout", p 324-331 v 148 n 3, Proc Am Phil Soc, Sep 2004 [YouTube]. Greg Bear, "When Genes Go Walkabout", p 324-331 v 148 n 3, Proc Am Phil Soc, Sep 2004 [YouTube].
 Thanks indirectly, Charles Siebert, "Unintelligent Design," v 27 n 3, Discover, Mar 2006. Thanks indirectly, Charles Siebert, "Unintelligent Design," v 27 n 3, Discover, Mar 2006.
 2000 2000
 2006 2006
 2014 2014
 2015: more from Villarreal. 2015: more from Villarreal.
 Viruses and Other Gene Transfer Mechanisms is the main related CA webpage [ Viruses and Other Gene Transfer Mechanisms is the main related CA webpage [ What'sNEW about HGT What'sNEW about HGT  ]. ].
...viruses are the dominate biological entities of the biosphere and are the most numerous, diverse and dynamic genetic agents on Earth
— Luis P. Villarreal
 24 February 2006 24 February 2006
Retroposed genes have contributed to human evolution, according to three Swiss geneticists. These are genes that have been reverse-transcribed from an edited RNA transcript, not copied by ordinary replication of DNA. They were once thought to be functionless, because they usually lack the regulatory elements needed to promote their own transcription. But these researchers write, "Our results suggest that retrocopy transcription is not rare...."The Swiss team first identified 3,950 retrocopies in the human genome. Of these, 575 have no disruptions that would prevent them from functioning, and these ones are almost twice as likely to be expressed as the disrupted ones. From here they estimate that the human genome contains at least 117 "bona fide retrogenes." Among their observations —
- "An overview of the most highly transcribed retrocopies... also provides compelling evidence for the correlation of transcription and functionality."
- "The top 50 transcribed retrocopies show significantly lower KA/KS values... than the remaining copies..., suggesting that they were predominantly shaped by purifying selection throughout their evolution."
- The retrogenes may rely on the regulatory mechanisms of neighboring genes, or acquire their own promoters by known methods. Some of the unexpressed ones may become expressed only after acquiring promoters. (These would be overlooked in the current study.)
- "Although selection generally acted against the insertion and high transcription of retrocopies located inside other genes, it favored the emergence of a substantial number of new genes with diverse gene structures and functions during the evolution of the human genome."
- "...We identified 27 other retrocopies transcribed together with additional exons. Our analyses show that these copies ...acquired new exons/introns de novo. ...We find a striking overrepresentation of such cases among highly expressed copies. For example, we identified 17 exon acquisition cases among the 50 most highly transcribed retrocopies (21 among the top 100), a > 4,200% excess relative to the 0.4 cases expected based on the overall data.... Several additional lines of evidence revealed that most of these retrocopies are not only highly expressed but represent bona fide genes."
- "...These analyses suggest that approximately one retrogene per million years has emerged on the primate lineage leading to humans," they write in an earlier (PLoS) article.
In the theory we promote, darwinian evolution can optimize and perfect genetic programs that are already available, but new programs must be acquired in a genetically open system. The story of retrogenes, and especially the examples of exon acquisition, are consistent with this picture, we think.
 Nicolas Vinckenbosch, Isabelle Dupanloup and Henrik Kaessmann, "Evolutionary fate of retroposed gene copies in the human genome" [abstract], 10.1073/pnas.0511307103, Proc. Natl. Acad. Sci. USA, online 21 Feb 2006. Nicolas Vinckenbosch, Isabelle Dupanloup and Henrik Kaessmann, "Evolutionary fate of retroposed gene copies in the human genome" [abstract], 10.1073/pnas.0511307103, Proc. Natl. Acad. Sci. USA, online 21 Feb 2006.
 Ana Claudia Marques et al., "Emergence of Young Human Genes after a Burst of Retroposition in Primates" [text], 10.1371/journal.pbio.0030357, e357 v3 n11 PLoS Biology, Nov 2005. Ana Claudia Marques et al., "Emergence of Young Human Genes after a Burst of Retroposition in Primates" [text], 10.1371/journal.pbio.0030357, e357 v3 n11 PLoS Biology, Nov 2005.
 Retroposition defined in MedTerms Medical Dictionary. Retroposition defined in MedTerms Medical Dictionary.
 Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ What'sNEW about HGT What'sNEW about HGT  ]. ].
 Human Genome Search... is a related CA webpage. Human Genome Search... is a related CA webpage.
 Robust Software Management (new, under construction) discusses other surprising mechanisms. Robust Software Management (new, under construction) discusses other surprising mechanisms.
 19 February 2006 19 February 2006
| | 
Cambrian body plans |
Why has there has been so little change in major body plans since the Early Cambrian, while great changes have subsequently occurred within phyla and classes? Two US biologists observe, "Classic evolutionary theory, based on selection of small incremental changes, has sought explanations by extrapolation from observed patterns of adaptation. Macroevolutionary theories have largely invoked multi-level selection, among species and among clades. But neither class of explanation provides an explanation of evolution in terms of mechanistic changes in the genetic regulatory program for development of the body plan, where it must lie." They believe that handfuls of genes they call the "kernels" of gene regulatory networks "must have been assembled during the initial diversification of the Bilateria ...but ...they would have become refractory to subsequent change." All of this is unsurprising for cosmic ancestry.
 Eric H. Davidson and Douglas H. Erwin, "Gene Regulatory Networks and the Evolution of Animal Body Plans" [abstract], 10.1126/science.1113832, p 796-800 v 311, Science, 10 February 2006. Eric H. Davidson and Douglas H. Erwin, "Gene Regulatory Networks and the Evolution of Animal Body Plans" [abstract], 10.1126/science.1113832, p 796-800 v 311, Science, 10 February 2006.
 Basic Body Design, This Week in STKE, 14 February 2006. Basic Body Design, This Week in STKE, 14 February 2006.
 NeoDarwinism... is the main related CA webpage. NeoDarwinism... is the main related CA webpage.
 18 February 2006 18 February 2006
Splicesomal introns are found in the nuclear genomes of all characterized eukaryotes. Their sequences are more varied and often far longer than the introns that are found in bacterial and organellar genomes. Unlike the latter, they require a complex of hundreds of proteins to remove them from RNA transcripts. Two Harvard biologists who are experts on introns recently reviewed what is known and unknown about the splicesomal ones. Because we suspect that introns may have a role in the transfer and installation of genetic programs, we studied the review carefully. Among the issues discussed in it —
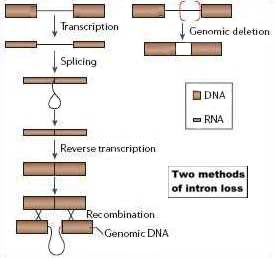
- How did the massive complexity of the spliceosome originate? "...Apart from a possible relationship between the spliceosome and the much simpler splicing mechanism of type II introns, no intermediate spliceosomal form has been found." The question is unanswered.
- How old are modern introns? Very old, the authors conclude. "...Only a minority [have] been inserted in the past hundreds of millions of years."
- Why are 25% of human introns at the exact same position as introns in the orthologous genes from a sunflower? The authors prefer to think that they were conserved from an ancient common ancestor, but parallel insertion could also be responsible.
- Did introns first appear early or late in the evolution of life on Earth? This question has been debated since the discovery of introns in the 1970s. The Harvard team favors the "Introns Early" theory, but they also write, "...Debate surrounds this issue. Studies have reached opposite conclusions. ...Contradictory results have been obtained.... Further comparative genomic studies will be necessary...."
- In the review's conclusion we read, "Another important question is why introns first arose. Despite 30 years of intense study, we are barely closer to answering this riddle." But if genetic programs are as old as life, and if they comprise mechanisms for transferring and inserting themselves into genomes, we think the riddle of introns makes more sense.
 Scott William Roy and Walter Gilbert, "The evolution of spliceosomal introns: patterns, puzzles and progress" [abstract], 10.1038/nrg1807, p 211-221 v 7, Nature Reviews Genetics, Mar (online Feb) 2006. Scott William Roy and Walter Gilbert, "The evolution of spliceosomal introns: patterns, puzzles and progress" [abstract], 10.1038/nrg1807, p 211-221 v 7, Nature Reviews Genetics, Mar (online Feb) 2006.
 Introns: a Mystery includes a definition and some background about introns, and a thumbnail picture of Walter Gilbert. Introns: a Mystery includes a definition and some background about introns, and a thumbnail picture of Walter Gilbert.
 16 February 2006 16 February 2006
 Ohio biology students must not analyze evolution. That's one way of interpreting a policy revision adopted 14 February by the Ohio Board of Education. The previous policy stated that public school students should be able to "describe how scientists continue to investigate and critically analyze aspects of evolutionary theory." The revision deletes this statement.
Ohio biology students must not analyze evolution. That's one way of interpreting a policy revision adopted 14 February by the Ohio Board of Education. The previous policy stated that public school students should be able to "describe how scientists continue to investigate and critically analyze aspects of evolutionary theory." The revision deletes this statement.
"The seemingly innocuous phrase 'critically analyze' is used by promoters of Intelligent Design (ID) to avoid making any anti-evolution references that could be construed as religious. The National Center for Science Education called the vote a 'stunning triumph' for Ohio students." Yes, the promoters of the previous policy may have had religious motives. And yes, religion should not be taught in science class. But science depends on critical analysis. If an analysis identifies flaws in existing theory, science should consequently be advanced, not diminished. Sadly, the false dilemma between darwinism and creationism gridlocks the subject of evolution. The Ohio Board's latest policy revision is better described as a stunning triumph for rigid darwinian dogma.
 Ohio Board Boots Anti-Evolution Policy, by Constance Holden, ScienceNow Daily News, 15 February 2006. Ohio Board Boots Anti-Evolution Policy, by Constance Holden, ScienceNow Daily News, 15 February 2006.
 Ohio Board of Education Votes 11-4..., Ohio Citizens for Science, 15 Feb 2006. Ohio Board of Education Votes 11-4..., Ohio Citizens for Science, 15 Feb 2006.
 Evolution vs Creationism is a related CA webpage. Evolution vs Creationism is a related CA webpage.
 ...A Third Alternative is a related CA webpage. ...A Third Alternative is a related CA webpage.
 14 February 2006 14 February 2006
Researchers evolve a complex genetic trait in the laboratory? This is how a project at Duke University has been announced. There a biology professor and a graduate student used artificial selection to produce a strain of tobacco hornworms whose color can be black or green, depending on incubation temperature. Normal tobacco hornworms found in nature are only green. But closely related tomato hornworms carry the temperature dependency, an example of "polyphenism." The biologists thought that the same polyphenism might be teased out of the tobacco hornworm.
 They started with a mutant tobacco hornworm, one that was black instead of the normal green. Thirteen generations of selective breeding with heat successfully yielded a strain which had the intended polyphenism. They name the evolutionary process "genetic accommodation."
They started with a mutant tobacco hornworm, one that was black instead of the normal green. Thirteen generations of selective breeding with heat successfully yielded a strain which had the intended polyphenism. They name the evolutionary process "genetic accommodation."
We believe that genetic programs for a feature like the polyphenism described here are noteworthy, and any demonstration that they were composed de novo by mutation and selection on unrelated DNA strands would be a monumental achievement. But no genetic sequences are reported in the study. Nevertheless the biologists suppose, "a mutation in a regulatory pathway such as a hormonal pathway changes the hormonal level to bring it closer to a threshold level that could be affected by environmental variation." If so, the programs were preexisting, and simply switched on. No one doubts that even a single point mutation can switch a regulatory program off or on. But this is very different from composing complex programs.
 Yuichiro Suzuki and H. Frederik Nijhout, "Evolution of a Polyphenism by Genetic Accommodation" [abstract], 10.1126/science.1118888, p 650-652, v 311, Science, 3 Feb 2006. Yuichiro Suzuki and H. Frederik Nijhout, "Evolution of a Polyphenism by Genetic Accommodation" [abstract], 10.1126/science.1118888, p 650-652, v 311, Science, 3 Feb 2006.
 Elizabeth Pennisi, "Hidden Genetic Variation Yields Caterpillar of a Different Color" [abstract], 10.1126/science.311.5761.591a, p 591, v 311, Science, 3 Feb 2006. Elizabeth Pennisi, "Hidden Genetic Variation Yields Caterpillar of a Different Color" [abstract], 10.1126/science.311.5761.591a, p 591, v 311, Science, 3 Feb 2006.
 "One Genome, Many Colors," [text], 10.1126/science.311.5761.573f, p 573, v 311, Science, 3 Feb 2006. "One Genome, Many Colors," [text], 10.1126/science.311.5761.573f, p 573, v 311, Science, 3 Feb 2006.
 Researchers Evolve A Complex Genetic Trait In The Laboratory, ScienceDaily.com, 13 Feb 2006. Researchers Evolve A Complex Genetic Trait In The Laboratory, ScienceDaily.com, 13 Feb 2006.
 Neodarwinism... is a related CA webpage. Neodarwinism... is a related CA webpage.
 Macroevolutionary Progress Redefined... is a related CA webpage. Macroevolutionary Progress Redefined... is a related CA webpage.
 Thanks, Jerry Chancellor. Thanks, Jerry Chancellor.
 13 February 2006 13 February 2006
Origin-of-life theory comes up short, according to David Deamer, emeritus professor of chemistry at the University of California, Santa Cruz. The clay scaffolding favored in some theories binds organic compounds so tightly that they cannot undergo further reactions, he says. (As usual, the "hardware" problem is not reduced, and the even more difficult "software" problem is ignored.) Deamer is among more than 200 leading scientists at an international meeting in London, 13-14 February, at the Royal Society, the UK national academy of science. The meeting will explore the latest thinking on the origin of life on Earth and other theories — including whether life arrived from space!

 Life on Earth 'unlikely to have emerged in volcanic springs', The Royal Society, 13 Feb 2006. Life on Earth 'unlikely to have emerged in volcanic springs', The Royal Society, 13 Feb 2006.
 Darwin's warm pond theory tested, by Rebecca Morelle, BBC News, 13 Feb 2006. Darwin's warm pond theory tested, by Rebecca Morelle, BBC News, 13 Feb 2006.
 Hot Soup Not So Tasty for Early Life, by Michael Schirber, ScienceNow Daily News, 15 Feb 2006. Hot Soup Not So Tasty for Early Life, by Michael Schirber, ScienceNow Daily News, 15 Feb 2006.
 The RNA World is the main CA webpage about origin-of-life theories. The RNA World is the main CA webpage about origin-of-life theories.
 Thanks, Klaas Dantuma and Sean Underwood. Thanks, Klaas Dantuma and Sean Underwood.
 12 February 2006 12 February 2006
 The Astrobiology Science Conference 2006 will be held 26-30 March in Washington DC, at the Ronald Reagan Building and International Trade Center. It is the fourth such conference; the first three were held at the NASA Ames Research Center, California. Astrobiology ...seeks to understand the origin and evolution of life on Earth, to determine if life exists elsewhere in the universe, and to predict the future of life on Earth and in the rest of the universe. To this end it relies on the diversity of disciplines and has inspired new meta-disciplines. Abstracts are solicited on all topics that span the enormous range of astrobiological themes. The meeting format will include a limited number of plenary talks that will complement a larger number of oral presentations in parallel thematic sessions.
The Astrobiology Science Conference 2006 will be held 26-30 March in Washington DC, at the Ronald Reagan Building and International Trade Center. It is the fourth such conference; the first three were held at the NASA Ames Research Center, California. Astrobiology ...seeks to understand the origin and evolution of life on Earth, to determine if life exists elsewhere in the universe, and to predict the future of life on Earth and in the rest of the universe. To this end it relies on the diversity of disciplines and has inspired new meta-disciplines. Abstracts are solicited on all topics that span the enormous range of astrobiological themes. The meeting format will include a limited number of plenary talks that will complement a larger number of oral presentations in parallel thematic sessions.
 AbSciCon 2006, 26-30 March, Washington DC. AbSciCon 2006, 26-30 March, Washington DC.
 10 February 2006 10 February 2006
 The UK Astrobiology Conference will be held at the University of Kent, 18-21 April. "The conference will cover all aspects of research related to astrobiology, including (but not exclusively): Astrobiology Technology, Mars, ExoMars, Panspermia, Microbial Colonies, Niche Environments, Extremophiles, Arctic and Antarctic Studies, Meteorites, Geomicrobiology, Prebiotic Climates, Human Life in Space, Planetary Protection/Contamination, Development of Life-forms in Other Environments, Origin of Life, Habitable Zones, Terrestrial Type Extra Solar Planets, Education and Outreach."
The UK Astrobiology Conference will be held at the University of Kent, 18-21 April. "The conference will cover all aspects of research related to astrobiology, including (but not exclusively): Astrobiology Technology, Mars, ExoMars, Panspermia, Microbial Colonies, Niche Environments, Extremophiles, Arctic and Antarctic Studies, Meteorites, Geomicrobiology, Prebiotic Climates, Human Life in Space, Planetary Protection/Contamination, Development of Life-forms in Other Environments, Origin of Life, Habitable Zones, Terrestrial Type Extra Solar Planets, Education and Outreach."
 UKAC06 — The second conference of the Astrobiology Society of Britain, 18-21 Apr 2006. UKAC06 — The second conference of the Astrobiology Society of Britain, 18-21 Apr 2006.
 9 February 2006 9 February 2006
 More evidence for past life on Mars comes from a new examination of the Nakhla meteorite which fell in Egypt, in 1911. Freshly fractured samples of it contain carbon structures that resemble features etched by microbes in volcanic glass on Earth. Carbon isotopes confirm that the inclusions are not terrestrial. Members of the Astrobiology Group at NASA's Johnson Space Center will present the finding in two papers at the 37th Lunar and Planetary Science Conference in Houston next month.
More evidence for past life on Mars comes from a new examination of the Nakhla meteorite which fell in Egypt, in 1911. Freshly fractured samples of it contain carbon structures that resemble features etched by microbes in volcanic glass on Earth. Carbon isotopes confirm that the inclusions are not terrestrial. Members of the Astrobiology Group at NASA's Johnson Space Center will present the finding in two papers at the 37th Lunar and Planetary Science Conference in Houston next month.

 D.S. McKay et al., "Observation and Analysis of in situ Carbonaceous Matter in Naklah: Part I" [2251.pdf], Lunar and Planetary Science XXXVII, 13-17 Mar 2006. D.S. McKay et al., "Observation and Analysis of in situ Carbonaceous Matter in Naklah: Part I" [2251.pdf], Lunar and Planetary Science XXXVII, 13-17 Mar 2006.
 E.K. Gibson et al., "Observation and Analysis of in situ Carbonaceous Matter in Naklah: Part II" [2039.pdf], Lunar and Planetary Science XXXVII, 13-17 Mar 2006. E.K. Gibson et al., "Observation and Analysis of in situ Carbonaceous Matter in Naklah: Part II" [2039.pdf], Lunar and Planetary Science XXXVII, 13-17 Mar 2006.
 The 37th Lunar and Planetary Science Conference, Houston TX, 13-17 March 2006. The 37th Lunar and Planetary Science Conference, Houston TX, 13-17 March 2006.
 New Viewing Technique Bolsters Case For Life On Mars, MarsDaily, 8 Feb 2006. New Viewing Technique Bolsters Case For Life On Mars, MarsDaily, 8 Feb 2006.
 Space rock re-opens Mars debate, by Paul Rincon, BBC News, 8 Feb 2006. Space rock re-opens Mars debate, by Paul Rincon, BBC News, 8 Feb 2006.
 Fresh Evidence for Life on Mars?, by David Biello, Scientific American, 9 Feb 2006. Fresh Evidence for Life on Mars?, by David Biello, Scientific American, 9 Feb 2006.
 Mars meteorite similar to bacteria-etched earth rocks, by Martin Fisk, EurekAlert!, 23 Mar 2006. Mars meteorite similar to bacteria-etched earth rocks, by Martin Fisk, EurekAlert!, 23 Mar 2006.
 Life on Mars! is a related CA webpage with a separate What'sNEW section about meteorites from Mars: ALH84001. Life on Mars! is a related CA webpage with a separate What'sNEW section about meteorites from Mars: ALH84001.
 Thanks, Jerry Chancellor and Jim Galasyn. Thanks, Jerry Chancellor and Jim Galasyn.
 9 February 2006 9 February 2006
A cell requires 430 genes according to a new study by the Synthetic Biology Group at the J. Craig Venter Institute. This is more than the 265-350 estimated by Venter's team in 1999. Starting with a bacterium that has "the smallest genome of any organism that can be grown in pure culture," Mycoplasma genitalium, "global transposon mutagenesis" was used to identify dispensable genes. The team concludes that even this minimal bacterium requires at least 387 protein-coding and 43 RNA-coding genes. Surprisingly, nearly a third of the necessary proteins have unknown functions. A chart from the study suggest how complex the metabolic pathways of even the simplest cell would be.

Metabolic pathways and substrate transport mechanisms encoded by M. genitalium. Metabolic products are colored red, and mycoplasma proteins are black. White letters on black boxes mark nonessential functions or proteins based on our current gene disruption study.... |
 John I. Glass et al., "Essential genes of a minimal bacterium" [abstract | pdf], doi:10.1073/pnas.0510013103, p 425-430 v 103, Proc. Natl. Acad. Sci. USA, 10 Jan (online 3 Jan) 2006. John I. Glass et al., "Essential genes of a minimal bacterium" [abstract | pdf], doi:10.1073/pnas.0510013103, p 425-430 v 103, Proc. Natl. Acad. Sci. USA, 10 Jan (online 3 Jan) 2006.
 "Absolute minimum," p 247 v 439, Nature, 19 Jan 2006. "Absolute minimum," p 247 v 439, Nature, 19 Jan 2006.
 What Is Life? explains that the simplest life is not simple. What Is Life? explains that the simplest life is not simple.
 Venter endorses panspermia, What'sNEW, 20 Jan 2005. Venter endorses panspermia, What'sNEW, 20 Jan 2005.

 7 February 2006 7 February 2006
Gerda Horneck of the German Aerospace Center has spent her career studying panspermia. In an interview she also discusses the effects of space radiation on life, the dangers of forward- and back-contamination, and issues that human astronauts would face.
 An interview with Gerda Horneck, Astrobiology Magazine, 6 Feb 2006. An interview with Gerda Horneck, Astrobiology Magazine, 6 Feb 2006.
 Introduction... is a related CA webpage. Introduction... is a related CA webpage.
 Thanks, Larry Klaes. Thanks, Larry Klaes.
 09 Jan 2002: Bacteria sent into space survive. 09 Jan 2002: Bacteria sent into space survive.
 2 February 2006 2 February 2006
| |
 Green = unique SMaseD plug with 7 conserved residues. Gray = strands and selected chains of the GDPD barrel. Green = unique SMaseD plug with 7 conserved residues. Gray = strands and selected chains of the GDPD barrel. |
A toxin found only in spiders and bacteria was probably installed by gene transfer, according to two US biologists. In the animal kingdom, a poisonous enzyme called sphingomyelinase D (SMaseD) is found only in the venom of certain spiders like the brown recluse. Elsewhere, it confers toxicity on strains of bacteria that infect farm animals. No other examples are known. "How does it come to pass that two similar, medically important protein toxins, putatively sharing a common evolutionary origin, are found in two very dissimilar types of organism but nowhere else?"The SMaseD enzyme shares homology with a ubiquitous domain family called GDPD that has a barrel shape. But in the GDPD family only SMaseD is toxic, and only it has a tail with seven perfectly conserved residues that plug the barrel. The US biologists wonder if "shared evolutionary pressure to gird the ...barrel sructure" could have caused the two distant species to stumble upon the same sequence of amino acids for a solution. No, many other sequences could plug the barrel, they observe. "Given the extraordinarily distant relationship between [the spiders and the bacteria], and the paucity of similar proteins in other organisms, LGT [Lateral Gene Transfer] is the most reasonable explanation for similarities between these toxins." The researchers even wonder if the spiders may have eaten the barnyard flies that vector the bacteria. By whatever means, two distant genomes acquired closely related genetic algorithms that have no obvious predecessor. The resulting phenotypic effects were important. This sounds like cosmic ancestry to us.
 Matthew H. J. Cordes and Greta J. Binford, "Lateral gene transfer of a dermonecrotic toxin between spiders and bacteria" [abstract], 10.1093/bioinformatics/bti811, p 264-268 v 22 n 3, Bioinformatics, 1 Feb 2006 (online 6 Dec 2005). Matthew H. J. Cordes and Greta J. Binford, "Lateral gene transfer of a dermonecrotic toxin between spiders and bacteria" [abstract], 10.1093/bioinformatics/bti811, p 264-268 v 22 n 3, Bioinformatics, 1 Feb 2006 (online 6 Dec 2005).
 Evolution mystery: Spider venom and bacteria share same toxin, by Tania Thompson, EurekAlert!, 1 Feb 2006. Evolution mystery: Spider venom and bacteria share same toxin, by Tania Thompson, EurekAlert!, 1 Feb 2006.
 Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ What'sNEW about HGT What'sNEW about HGT  ]. ].
 30 January 2006 30 January 2006
More survivors of the Columbia disintegration have been found by biologists at Texas State University, San Marcos. In a planned experiment three species of microbes sealed inside plastic containers within aluminum cases were launched on the mission, 16 January 2003. When Columbia burned up in the high atmosphere during reentry, 1 February 2006, one of the containers crashed onto a Texas parking lot. The heat or 2500g impact forces had killed the original microbes in it, but a fourth species not listed on the mission manifest, a slow-growing, heat-tolerant soil bacterium, survived. "Because the containers' seals were intact, the stowaway organism [must have been] an airborne contaminant that got into the container during assembly before the launch."
 "Their survival is one link in a chain of evidence needed to prove the plausibility of panspermia," said Bruce Runnegar, science director at NASA's Astrobiology Institute. "But it's a very encouraging result.... I think all reasonable scientists regard the idea as plausible." Earlier, in November 2005, a NASA team reported that nematodes also survived in canisters found among Columbia wreckage that fell near San Augustine and Bronson, Texas.
"Their survival is one link in a chain of evidence needed to prove the plausibility of panspermia," said Bruce Runnegar, science director at NASA's Astrobiology Institute. "But it's a very encouraging result.... I think all reasonable scientists regard the idea as plausible." Earlier, in November 2005, a NASA team reported that nematodes also survived in canisters found among Columbia wreckage that fell near San Augustine and Bronson, Texas.
 Prof says life seeds survived Columbia, by Roger Croteau, San Antonio Express-News, 5 Mar 2006. Prof says life seeds survived Columbia, by Roger Croteau, San Antonio Express-News, 5 Mar 2006.
 Keeping cool while hitchhiking from space, by Frank D. Roylance, BaltimoreSun.com, 27 Jan 2006. Keeping cool while hitchhiking from space, by Frank D. Roylance, BaltimoreSun.com, 27 Jan 2006.
 R.J.C. McLean et al., "Microbial survival in space shuttle crash," Icarus, in press 2006. R.J.C. McLean et al., "Microbial survival in space shuttle crash," Icarus, in press 2006.
 Nathaniel J. Szewczyk, et al., "Caenorhabditis elegans Survives Atmospheric Breakup of STS-107, Space Shuttle Columbia" [PDF], v 5 n 6, Astrobiology, Nov 2005. Nathaniel J. Szewczyk, et al., "Caenorhabditis elegans Survives Atmospheric Breakup of STS-107, Space Shuttle Columbia" [PDF], v 5 n 6, Astrobiology, Nov 2005.
 Introduction... is a related CA webpage. Introduction... is a related CA webpage.
 Bacteria... is a related CA webpage. Bacteria... is a related CA webpage.
 Thanks, Robert Cobb. Thanks, Robert Cobb.
 28 January 2006 28 January 2006
 Important aspects of the history of life are replicable and predictable, according to a well-documented analysis by paleoecologist Geerat J. Vermeij of the University of California, Davis. His conclusion contradicts the standard darwinian line, "future states cannot be predicted even one step away from the present."
Important aspects of the history of life are replicable and predictable, according to a well-documented analysis by paleoecologist Geerat J. Vermeij of the University of California, Davis. His conclusion contradicts the standard darwinian line, "future states cannot be predicted even one step away from the present."
 Vermeij analyzes 23 innovations usually considered unique (left), and 55 repeated innovations like nitrogen fixation, multicellularity, plant leaves, land-plant vines, mineralized skeletons, image-forming eyes, tetrapod wings, vertebrate gliding, vertebrate teeth and the vertebrate placenta (an eye-opening list!) Three-fifths of the former, but ony 16% of the known repeated ones, first occurred before 543 million years ago. Were unique innovations simply more common earlier? Vermeij decides no, they only appear so because of information loss — some ancient lineages have left no descendants to sample, and others cannot be discriminated after so much accumulated genetic change. He concludes, "few, if any, innovations are truly unique." Logically, therefore, "similar phenotypes ...should emerge wherever conditions suitable for life exist."
Vermeij analyzes 23 innovations usually considered unique (left), and 55 repeated innovations like nitrogen fixation, multicellularity, plant leaves, land-plant vines, mineralized skeletons, image-forming eyes, tetrapod wings, vertebrate gliding, vertebrate teeth and the vertebrate placenta (an eye-opening list!) Three-fifths of the former, but ony 16% of the known repeated ones, first occurred before 543 million years ago. Were unique innovations simply more common earlier? Vermeij decides no, they only appear so because of information loss — some ancient lineages have left no descendants to sample, and others cannot be discriminated after so much accumulated genetic change. He concludes, "few, if any, innovations are truly unique." Logically, therefore, "similar phenotypes ...should emerge wherever conditions suitable for life exist."
Beyond the data he justifies his conclusion with darwinian orthodoxy. For example, "The conditions necessary for the formation of true cells should have been common and widespread, making a unique origin [of life] highly unlikely." (So life on Earth probably originated lots of times? Howcome the process can't be demonstrated?) Vermeij acknowledges the importance of symbiosis, now also part of darwinian orthodoxy. And he thinks gene transfer has been rampant among prokaryotes, but is rarer for eukaryotes. (Does he know that only three or four successfully transferred genes per million years could furnish the estmated 300 genes separating humans from mice?) In our amended theory of evolution, life requires genetic programs that are preexisting, fairly ubiquitous and readily transferred. If so, yes, evolutionary innovations would be replicable and predictable, and life elsewhere would resemble Earthly life.
 Geerat J. Vermeij, "Historical contingency and the purported uniqueness of evolutionary innovations" [abstract], 10.1073/pnas.0508724103, p 1804-1809 v 103, Proc. Natl. Acad. Sci. USA, 7 Feb (online 27 Jan) 2006. Geerat J. Vermeij, "Historical contingency and the purported uniqueness of evolutionary innovations" [abstract], 10.1073/pnas.0508724103, p 1804-1809 v 103, Proc. Natl. Acad. Sci. USA, 7 Feb (online 27 Jan) 2006.
 Life, the Remake, UC Davis News Service, 24 Feb 2006. Life, the Remake, UC Davis News Service, 24 Feb 2006.
 Geerat J. Vermeij, Department of Geology, The University of California, Davis. Geerat J. Vermeij, Department of Geology, The University of California, Davis.
 Neo-Darwinsm..., a related CA webpage, has a section on Convergent Evolution. Neo-Darwinsm..., a related CA webpage, has a section on Convergent Evolution.
 Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ What'sNEW about HGT What'sNEW about HGT  ]. ].
 Macroevolutionary Progress ...Without Gene Transfer? is a related CA webpage. Macroevolutionary Progress ...Without Gene Transfer? is a related CA webpage.
 Ron McGhee replies in agreement, 1 Feb 2006. Ron McGhee replies in agreement, 1 Feb 2006.

 18 January 2006 18 January 2006
We hypothesize that these 'jumping genomic segments' are part of an ongoing evolutionary process that results in a novel form of large-scale variation in human genomic DNA and contributes rapidly to primate gene evolution — Evan Eichler
 Evan Eichler, Associate Professor of Genome Sciences, The University of Washington. Evan Eichler, Associate Professor of Genome Sciences, The University of Washington.
 Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ Viruses and Other Gene Transfer Mechanisms is a related CA webpage [ What'sNEW about HGT What'sNEW about HGT  ]. ].
 Human Genome Search... is a related CA webpage. Human Genome Search... is a related CA webpage.
 15 January 2006 15 January 2006
 Stardust landed safely (softly) early this morning. Hooray!
Stardust landed safely (softly) early this morning. Hooray!
 NASA's Comet Tale Draws to a Successful Close in Utah Desert, NASA News Release: 2006-009, 15 Jan 2006. NASA's Comet Tale Draws to a Successful Close in Utah Desert, NASA News Release: 2006-009, 15 Jan 2006.
 Stardust parachutes to soft landing in Utah with dust samples from comet, University of Washington News Office, 15 Jan 2006. Stardust parachutes to soft landing in Utah with dust samples from comet, University of Washington News Office, 15 Jan 2006.
 Magnet Lab to Analyze Stardust Mission's Comet Dust, Florida State University Communications, 13 Jan 2006. Magnet Lab to Analyze Stardust Mission's Comet Dust, Florida State University Communications, 13 Jan 2006.
 Comet dust delivered to Earth, by Mark Peplow, News@Nature.com, 16 Jan 2006. Comet dust delivered to Earth, by Mark Peplow, News@Nature.com, 16 Jan 2006.
 Stardust Capsule Returns to Earth, Astronomy Picture of the Day, 16 Jan 2006. Stardust Capsule Returns to Earth, Astronomy Picture of the Day, 16 Jan 2006.
 7 January 2006 7 January 2006
Stardust will return to Earth for a soft landing at the U.S. Air Force Utah Test and Training Range early in the morning, 15 January. A planned steering maneuver went flawlessly Thursday. The capsule will bring particles gathered during its close rendezvous with comet Wild 2 ("vilt 2"), 2 January 2004. "Scientists believe in-depth terrestrial analysis of cometary samples will reveal much not just about comets but about the earliest history of the solar system."
 Stardust Mission Status Report, NASA, 5 Jan 2006. Stardust Mission Status Report, NASA, 5 Jan 2006.
 Comets... is a related CA webpage. Comets... is a related CA webpage.
 ...Interstellar Dust is a related CA webpage. ...Interstellar Dust is a related CA webpage.
 Comet Rendezvous is a related section of the webpage, "Can The Theory Be Tested?", with links to earlier articles about Stardust. Comet Rendezvous is a related section of the webpage, "Can The Theory Be Tested?", with links to earlier articles about Stardust.
 6 January 2006 6 January 2006
| | 
SEM image of red rain cells under high magnification (8000x) |
A dust storm couldn't have caused the red rain of Kerala, according to the two scientists at Mahatma Gandhi University who originally researched the 2001 phenomenon on the west coast of southern India. In a new paper they note that a long dry spell would be needed for windborne dust to accumulate, but the first red rain followed closely after a normal rain. Also, its distribution was patchy, whereas dust from across the Arabian Sea would be evenly dispersed. Besides, the red particles are homogeneous and unlike typical desert dust. They appear biological in both form and chemistry, although they apparently lack DNA or RNA. At least 50,000 kg of the particles fell with the rain, most of it within the first two weeks of the phenomenon. The two investigators believe the particles came from fragments of a meteorite that burst in the atmosphere. Analysis continues.
 Godfrey Louis and A. Santhosh Kumar, "The Red Rain Phenomenon of Kerala and Its Possible Extraterrestrial Origin" [abstract | PDF: 18 pages], arXiv:astro-ph/0601022 v1 2 Jan 2006, accepted for publication in Astrophysics and Space Science, 1 Jan 2006. Godfrey Louis and A. Santhosh Kumar, "The Red Rain Phenomenon of Kerala and Its Possible Extraterrestrial Origin" [abstract | PDF: 18 pages], arXiv:astro-ph/0601022 v1 2 Jan 2006, accepted for publication in Astrophysics and Space Science, 1 Jan 2006.
 The red rain of Kerala — our initial What'sNEW report of the investigation, 23 Oct 2003. The red rain of Kerala — our initial What'sNEW report of the investigation, 23 Oct 2003.
 A dust storm caused the red rain...? — What'sNEW, 24 Dec 2005. A dust storm caused the red rain...? — What'sNEW, 24 Dec 2005.
 Thanks, Larry Klaes, Benny Peiser, Richard Hoover, and Thanks, Larry Klaes, Benny Peiser, Richard Hoover, and  Ian Goddard for your reply of 28 Mar 2006. Ian Goddard for your reply of 28 Mar 2006.
|


 Ohio biology students must not analyze evolution. That's one way of interpreting a policy revision adopted 14 February by the Ohio Board of Education. The previous policy stated that public school students should be able to "describe how scientists continue to investigate and critically analyze aspects of evolutionary theory." The revision deletes this statement.
Ohio biology students must not analyze evolution. That's one way of interpreting a policy revision adopted 14 February by the Ohio Board of Education. The previous policy stated that public school students should be able to "describe how scientists continue to investigate and critically analyze aspects of evolutionary theory." The revision deletes this statement.

 They started with a mutant tobacco hornworm, one that was black instead of the normal green. Thirteen generations of selective breeding with heat successfully yielded a strain which had the intended polyphenism. They name the evolutionary process "genetic accommodation."
They started with a mutant tobacco hornworm, one that was black instead of the normal green. Thirteen generations of selective breeding with heat successfully yielded a strain which had the intended polyphenism. They name the evolutionary process "genetic accommodation."

 The UK Astrobiology Conference will be held at the University of Kent, 18-21 April. "The conference will cover all aspects of research related to astrobiology, including (but not exclusively): Astrobiology Technology, Mars, ExoMars, Panspermia, Microbial Colonies, Niche Environments, Extremophiles, Arctic and Antarctic Studies, Meteorites, Geomicrobiology, Prebiotic Climates, Human Life in Space, Planetary Protection/Contamination, Development of Life-forms in Other Environments, Origin of Life, Habitable Zones, Terrestrial Type Extra Solar Planets, Education and Outreach."
The UK Astrobiology Conference will be held at the University of Kent, 18-21 April. "The conference will cover all aspects of research related to astrobiology, including (but not exclusively): Astrobiology Technology, Mars, ExoMars, Panspermia, Microbial Colonies, Niche Environments, Extremophiles, Arctic and Antarctic Studies, Meteorites, Geomicrobiology, Prebiotic Climates, Human Life in Space, Planetary Protection/Contamination, Development of Life-forms in Other Environments, Origin of Life, Habitable Zones, Terrestrial Type Extra Solar Planets, Education and Outreach." More evidence for past life on Mars comes from a new examination of the Nakhla meteorite which fell in Egypt, in 1911. Freshly fractured samples of it contain carbon structures that resemble features etched by microbes in volcanic glass on Earth. Carbon isotopes confirm that the inclusions are not terrestrial. Members of the Astrobiology Group at NASA's Johnson Space Center will present the finding in two papers at the 37th Lunar and Planetary Science Conference in Houston next month.
More evidence for past life on Mars comes from a new examination of the Nakhla meteorite which fell in Egypt, in 1911. Freshly fractured samples of it contain carbon structures that resemble features etched by microbes in volcanic glass on Earth. Carbon isotopes confirm that the inclusions are not terrestrial. Members of the Astrobiology Group at NASA's Johnson Space Center will present the finding in two papers at the 37th Lunar and Planetary Science Conference in Houston next month.



 "Their survival is one link in a chain of evidence needed to prove the plausibility of panspermia," said Bruce Runnegar, science director at NASA's Astrobiology Institute. "But it's a very encouraging result.... I think all reasonable scientists regard the idea as plausible." Earlier, in November 2005, a NASA team reported that nematodes also survived in canisters found among Columbia wreckage that fell near San Augustine and Bronson, Texas.
"Their survival is one link in a chain of evidence needed to prove the plausibility of panspermia," said Bruce Runnegar, science director at NASA's Astrobiology Institute. "But it's a very encouraging result.... I think all reasonable scientists regard the idea as plausible." Earlier, in November 2005, a NASA team reported that nematodes also survived in canisters found among Columbia wreckage that fell near San Augustine and Bronson, Texas. Important aspects of the history of life are replicable and predictable, according to a well-documented analysis by paleoecologist Geerat J. Vermeij of the University of California, Davis. His conclusion contradicts the standard darwinian line, "future states cannot be predicted even one step away from the present."
Important aspects of the history of life are replicable and predictable, according to a well-documented analysis by paleoecologist Geerat J. Vermeij of the University of California, Davis. His conclusion contradicts the standard darwinian line, "future states cannot be predicted even one step away from the present."
 Vermeij analyzes 23 innovations usually considered unique (left), and 55 repeated innovations like nitrogen fixation, multicellularity, plant leaves, land-plant vines, mineralized skeletons, image-forming eyes, tetrapod wings, vertebrate gliding, vertebrate teeth and the vertebrate placenta (an eye-opening list!) Three-fifths of the former, but ony 16% of the known repeated ones, first occurred before 543 million years ago. Were unique innovations simply more common earlier? Vermeij decides no, they only appear so because of information loss — some ancient lineages have left no descendants to sample, and others cannot be discriminated after so much accumulated genetic change. He concludes, "few, if any, innovations are truly unique." Logically, therefore, "similar phenotypes ...should emerge wherever conditions suitable for life exist."
Vermeij analyzes 23 innovations usually considered unique (left), and 55 repeated innovations like nitrogen fixation, multicellularity, plant leaves, land-plant vines, mineralized skeletons, image-forming eyes, tetrapod wings, vertebrate gliding, vertebrate teeth and the vertebrate placenta (an eye-opening list!) Three-fifths of the former, but ony 16% of the known repeated ones, first occurred before 543 million years ago. Were unique innovations simply more common earlier? Vermeij decides no, they only appear so because of information loss — some ancient lineages have left no descendants to sample, and others cannot be discriminated after so much accumulated genetic change. He concludes, "few, if any, innovations are truly unique." Logically, therefore, "similar phenotypes ...should emerge wherever conditions suitable for life exist."

 Stardust landed safely (softly) early this morning. Hooray!
Stardust landed safely (softly) early this morning. Hooray!