What Is Life? Updates since 1999
I am not going to answer this question — J. B. S. Haldane (1)
The units of life are cells — Lynn Margulis (2)
Life Is Cells
People like to say, as if it were obvious, that life is hard to define. This is misleading. Life has properties that clearly distinguish it from everything else. First, every living thing is cellular. In other words, it is either a single-celled creature or a creature composed of many cells. Every cell is bounded by its own outer membrane and contains a full set of instructions necessary for its operation and reproduction. Furthermore, every cell uses the same operating system: "DNA makes RNA makes protein." DNA is a long complex molecule that contains the cell's instructions. It is transcribed into RNA, another long complex molecule similar to DNA; and then the RNA transcript is translated into protein. There are hundreds of billions of different proteins used by living things (3), but all of them are made from the same twenty amino acids, the "building blocks of life."
Other Properties of Life
 Living things reproduce themselves. Either individually or in sexual pairs, they have both the encoded instructions and the machinery necessary for self-reproduction. (Some creatures cannot reproduce, but every creature comes from reproduction.) Periodic crystals like sodium chloride (table salt) also undergo a kind of self-reproduction. In crystals however, the "instructions" are much simpler, they are not encoded, and they are not different from the "machinery." Living things reproduce themselves. Either individually or in sexual pairs, they have both the encoded instructions and the machinery necessary for self-reproduction. (Some creatures cannot reproduce, but every creature comes from reproduction.) Periodic crystals like sodium chloride (table salt) also undergo a kind of self-reproduction. In crystals however, the "instructions" are much simpler, they are not encoded, and they are not different from the "machinery."
 Life uses processes collectively called metabolism to convert materials and energy for its needs. Metabolism creates waste products. When metabolism ceases with no prospect of starting again, we call it death. Machines also convert materials and energy for their needs, create waste, and could be said to die. Life uses processes collectively called metabolism to convert materials and energy for its needs. Metabolism creates waste products. When metabolism ceases with no prospect of starting again, we call it death. Machines also convert materials and energy for their needs, create waste, and could be said to die.
 Life undergoes evolution. Notably, simpler forms are succeeded by forms with greater organization. Cars evolve also, in their way. Computers do, too. And computers even contain their own encoded instruction sets. Life undergoes evolution. Notably, simpler forms are succeeded by forms with greater organization. Cars evolve also, in their way. Computers do, too. And computers even contain their own encoded instruction sets.
| What Is Motor Vehicle Traffic? It is tempting to say that motor vehicle traffic is simply the things that move along the streets and highways — cars and trucks. Of course buses and motorcycles should be included, although they are absent or prohibited on some streets. But what about wheelchairs, bicycles and skateboards — sometimes these are motorized. What about a trailer that is merely towed behind a tractor? What about a tire that happens to come off and roll a tenth of a mile? What about rocks that fall out of a dump truck and bounce and skid along the highway? As it turns out, motor vehicle traffic is quite difficult to define. But naturally it would be hard to draw a line between cars and trucks, and the skidding rocks from which they must have evolved.
What About Airplanes? Suppose an attendant at turnpike toll-booth is ignorant of airplanes, but on holiday, he hears and sees them overhead. Does he recognize another kind of motor vehicle traffic? |
These latter properties of life are sometimes used to make the point that life is hard to define. But nothing else has all of these latter properties except cellular life using life's DNA—RNA—protein operating system. Another kind of life, entirely different from ours, is conceivable, yes. But the only kind we have ever seen is the one we are part of here on Earth. As biologist and philosopher Harold J. Morowitz says, "The only life we know for certain is cellular..." (4).
Viruses and prions are not alive; they lie on the fringe of life. Viruses contain instructions encoded in DNA or RNA. (Prions don't.) Both are reproduced. Viruses certainly and prions probably can evolve. But neither can reproduce itself; each needs the machinery of a living cell to carry out its reproduction. Without a cell, viruses and prions are merely inert, complicated particles which do nothing. Do they make it hard to define life? No, just as trailers don't make it hard to define motor vehicle traffic. We know what motor vehicle traffic is. And we know what life is.
The concept of the gene as a symbolic representation of the organism – a code script – is a fundamental feature of the living world and must form the kernel of biological theory — Sydney Brenner, 2012 (4.2)
All the regularities of biology strike me as being exactly like the regularities of engineering — Daniel C. Dennett (4.5)
One analogy for a cell is a computer. Computers have coded instructions inside them called programs. The programs in computers are analogous to the genetic programming in the DNA within cells. DNA is subdivided into functional units called genes; these would correspond to files in the computer. A computer even has a metabolism: it consumes electrical energy and discharges heat.
The programs in cells and those in computers can both be 1) copied and 2) executed. Some of the proteins made when a genetic program is executed would loosely correspond to the computer's paper printout. But other proteins are more analogous to the computer's cabinetry or wiring. Of course, computers don't make their own cabinetry or wiring; the analogy is not perfect.
In fact, nothing about the computer is analogous to a cell's reproduction. A cell can make a complete copy of itself; it contains the complete instructions (programs) and the cellular machinery (hardware) necessary to reproduce itself. A computer cannot make a copy of itself. It lacks the necssary machinery (but it may be able to reproduce its instruction set by "automatic full backup".) A computer that could reproduce itself would be more properly described as a self-reproducing robot. Such a thing is conceivable, but none exists on Earth today.
A multicelled creature is like a network of computers. It requires parallel computer architecture on a huge scale to operate multicelled creatures such as mammals with millions of millions of cells, all working in harmony, each doing its task. The nervous system and the hormonal system are two important networking systems used by mammals.
Changing the way a computer works requires new programs. Sometimes one can simply insert a disc into a slot: the computer recognizes the disc, accepts its new code, and uses it. Other times, reprogramming a computer is more trouble. The new software may have "bugs"; it may not be compatible with the existing software; additional software patches may be needed; it may introduce a computer virus; or it may cause everything to crash without explanation.
Biological evolution happens when cells are reprogrammed. Somehow, new genetic programs are installed and activated. How does new genetic software get installed and activated? And where does it come from? These are questions that Cosmic Ancestry attempts to answer.
The Two Kinds of Cells
There are two kinds of cells. You might guess the two are plant and animal cells. This distinction, however, is even more profound. The two kinds are prokaryotes and eukaryotes. (All plant and animal cells are eukaryotic.)
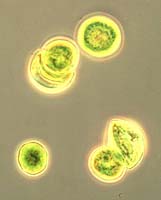 bacteria / Susan M. Barns bacteria / Susan M. Barns |
Prokaryotes are smaller and simpler than eukaryotic cells. They have no cell nucleus. They can multiply faster than eukaryotic cells, mainly for two reasons: 1) They have shorter genetic instructions to be replicated; and 2) The replication process goes about ten times as fast. Prokaryotes don't combine and specialize to form multicelled creatures. Prokaryotes are also called bacteria. They come in a wide variety of types; their diversity is much greater than that of eukaryotes.
Prokaryotes were here first, appearing very soon after Earth had cooled enough for life to survive. The oldest rocks that could contain recognizable fossils contain evidence of domelike structures left by colonies of cyanobacteria and other bacteria. Even older rocks contain chemical evidence that bacterial metabolism was under way (5).
Prokaryotes are divided into two major subkingdoms: eubacteria and archaebacteria. Eubacteria, or "true bacteria", are more familiar and ubiquitous, thriving in soil, water, our own mouths, etc. Archaebacteria differ from eubacteria in some basic ways. For example, their ribosomes (nanoscale protein factories) have a different shape. In fact, archaebacteria are in some ways more similar to eukaryotes than to eubacteria. Biologists now think, based on the reconstruction of genetic "trees," that archaebacteria are the oldest kind of cell. Today some biologists maintain that archaebacteria constitute a third domain of life which could be called simply archaea (6-9).
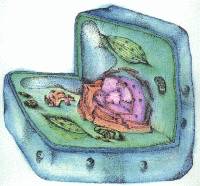 eukaryote / The ESG Biology Hypertextbook eukaryote / The ESG Biology Hypertextbook |
Eukaryotic cells are much more complicated than prokaryotic cells. The eukaryotic cell has a differentiated nucleus enclosed in a nuclear membrane. It usually has two whole copies of the genome, so in computer terms the eukaryotic cell has a backup copy of its programs. Outside of its nucleus, the eukaryotic cell has an array of complex subunits that are essential to it. Two of the subunits, mitochondria and chloroplasts, have their own DNA. These two subunits enable eukaryotic cells that contain them to conduct respiration and photosynthesis, respectively. And a eukaryotic cell has an internal "cytoskeleton" that supports, transports and rearranges its structure and elements. With these features, eukaryotic cells are able to constitute multicelled animals and plants. Thus, eukaryotes are able to acquire much more complex organs and systems than prokaryotes. If life has existed on Earth for almost four billion years, the consensus is that eukaryotes first appeared just after the halfway point, maybe 1.7 billion years ago.
Returning to the computer analogy, the relationship between prokaryotes and eukaryotes is like the relationship between handheld calculators and desktop personal computers. Both kinds of cells come in a broad range of sizes, but prokaryotes are, on average, about an order of magnitude smaller, like handheld calculators. And they come in a wide variety, each with a narrow special purpose. Consider scientific calculators, inventory scanners, GPS units, cellphones, cordless phones, pagers, beepers, walkie-talkies, PDAs, TV remote controllers, keyless entry buttons, Gameboys, Walkmans, iPods, guitar tuners, electronic or medical diagnostic kits, digital cameras, smoke detectors, portable radios, digital thermometers and cordless shavers. Like eukaryotes, personal computers have greater memory capacity, have more complicated structure, and can be networked (eukaryotes form multicelled creatures).
The size of a cell's genome can be compared to the amount of programming stored in a computer, using the equation, 4 nucleotides = 8 bits = 1 byte. The simplest prokaryotic cell would correspond to a handheld calculator with about 200 kilobytes of stored programs; the E. coli bacterium would correspond to a handheld calculator with about 1.2 megabytes of stored programs. Among eukaryotic cells, counting the backup copy of the genome and the "silent" DNA, a yeast cell would correspond to a personal computer with 12 megabytes of program storage capacity; a human cell corresponds to a personal computer with 1.5 gigabytes of program storage capacity. And the human body would correspond to a computer network of a hundred trillion (10^14) or more such units.
Updates
We do not have to define "life" in an abstract way — Leslie E. Orgel (10)
I'm not sure that we need to bother about a precise definition — Denis Noble (11)
 "Bringing the genetically minimal cell to life on a computer in 4D," by Zane R. Thornburg, Andrew Maytin et al, Cell, 09 Mar 2026.
...a single 543 kbp circular chromosome ...of 493 genes. "Bringing the genetically minimal cell to life on a computer in 4D," by Zane R. Thornburg, Andrew Maytin et al, Cell, 09 Mar 2026.
...a single 543 kbp circular chromosome ...of 493 genes.
 "Symbiotic bacteria ...break record for smallest non-organelle genome...," by Krystal Kasal, Phys.Org, 20 Feb 2026; re: Anna Michalik et al, doi:10.1038/s41467-026-69238-x, Nature Communications, 07 Feb 2026.
...loss of genes related to cellular functions makes these bacteria highly dependent on their host, almost like organelles. "Symbiotic bacteria ...break record for smallest non-organelle genome...," by Krystal Kasal, Phys.Org, 20 Feb 2026; re: Anna Michalik et al, doi:10.1038/s41467-026-69238-x, Nature Communications, 07 Feb 2026.
...loss of genes related to cellular functions makes these bacteria highly dependent on their host, almost like organelles.
 "Fundamental constraints to the logic of living systems," by Ricard Solé et al., J. R. Soc. Interface Focus, 2024.
Even the simplest cells exhibit an extraordinary and diverse molecular complexity and common design principles. "Fundamental constraints to the logic of living systems," by Ricard Solé et al., J. R. Soc. Interface Focus, 2024.
Even the simplest cells exhibit an extraordinary and diverse molecular complexity and common design principles.
 16 Sep 2024: How did eukaryotic cells evolve? 16 Sep 2024: How did eukaryotic cells evolve?
 New astrobiology research predicts life 'as we don't know it', by Karin Valentine, ASU News, 28 Feb 2022; re: New astrobiology research predicts life 'as we don't know it', by Karin Valentine, ASU News, 28 Feb 2022; re:
 Scaling laws ...reveal a new kind of biochemical universality by Dylan C. Gagler et al., PNAS, 25 Feb 2022. Scaling laws ...reveal a new kind of biochemical universality by Dylan C. Gagler et al., PNAS, 25 Feb 2022.
 25 Feb 2022: A computer model of a minimal cell contains 493 genes. 25 Feb 2022: A computer model of a minimal cell contains 493 genes.
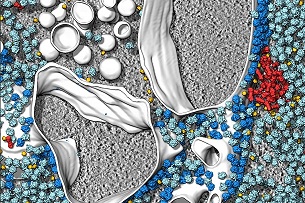
 ...cells – as never seen before by Diana Kwon, Nature, 26 Oct 2021 (see right). ...cells – as never seen before by Diana Kwon, Nature, 26 Oct 2021 (see right).
 20 Aug 2021: A new definition of life from Santa Fe Institute. 20 Aug 2021: A new definition of life from Santa Fe Institute.
 What Is Life?... by Carl Zimmer, Quanta, 09 Mar 2021. ...there are as many definitions of life as there are people trying to define it. What Is Life?... by Carl Zimmer, Quanta, 09 Mar 2021. ...there are as many definitions of life as there are people trying to define it.
 06 Sep 2020: What is Lyfe? 06 Sep 2020: What is Lyfe?
 ...life-like cells from scratch by Kendall Powell, Nature, 07 Nov 2018. ...could soon test the boundaries of life. ...life-like cells from scratch by Kendall Powell, Nature, 07 Nov 2018. ...could soon test the boundaries of life.
 Detecting Life On Other Worlds by Alex Tolley, intro by Paul Gilster, Centauri Dreams, 10 Aug 2018. Detecting Life On Other Worlds by Alex Tolley, intro by Paul Gilster, Centauri Dreams, 10 Aug 2018.
 "Life and life only: a radical alternative to life definitionism" by Carlos Mariscal and W. Ford Doolittle, doi:10.1007/s11229-018-1852-2, Synthese, 09 Jun 2018. "Life and life only: a radical alternative to life definitionism" by Carlos Mariscal and W. Ford Doolittle, doi:10.1007/s11229-018-1852-2, Synthese, 09 Jun 2018.
 06 Jan 2017: Either life developed here super-fast or it came full-on as DNA life from afar — Gary Ruvkun, SETG. 06 Jan 2017: Either life developed here super-fast or it came full-on as DNA life from afar — Gary Ruvkun, SETG.
 09 Dec 2016: The History and Philosophy of Origin Research (Tokyo conference report) 09 Dec 2016: The History and Philosophy of Origin Research (Tokyo conference report)
 Are viruses alive?.. by Eugene V. Koonin and Petro Starokadomskyy, do:10.1016/j.shpsc.2016.02.016, Studies in History and Philosophy of Biological and Biomedical Science, Oct 2016. Are viruses alive?.. by Eugene V. Koonin and Petro Starokadomskyy, do:10.1016/j.shpsc.2016.02.016, Studies in History and Philosophy of Biological and Biomedical Science, Oct 2016.
 26 Mar 2016: ...we set out to construct a minimal cellular genome.... Syn3.0 has a 531-kbp genome that encodes 473 gene products. 26 Mar 2016: ...we set out to construct a minimal cellular genome.... Syn3.0 has a 531-kbp genome that encodes 473 gene products.
 7 Aug 2015: A wide-ranging panel discussion...(link to video of ASU conference held 12 Feb 2011.) 7 Aug 2015: A wide-ranging panel discussion...(link to video of ASU conference held 12 Feb 2011.)
 Hanna K. E. Landenmark et al., "An Estimate of the Total DNA in the Biosphere," doi:10.1371/journal.pbio.1002168, PLoS Biolgy, 11 Jun 2015. "By analogy, it would require 10^21 computers with the mean storage capacity of the world's four most powerful supercomputers (Tianhe-2, Titan, Sequoia, and K computer) to store this information." Hanna K. E. Landenmark et al., "An Estimate of the Total DNA in the Biosphere," doi:10.1371/journal.pbio.1002168, PLoS Biolgy, 11 Jun 2015. "By analogy, it would require 10^21 computers with the mean storage capacity of the world's four most powerful supercomputers (Tianhe-2, Titan, Sequoia, and K computer) to store this information."
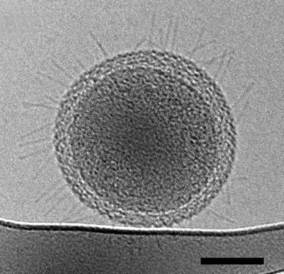
 First Detailed Microscopy Evidence of Bacteria at the Lower Size Limit of Life by Dan Krotz, Berkeley Lab (+Newswise), 27 Feb 2015. (See TEM image at right; scale bar = 100 nanometers.) First Detailed Microscopy Evidence of Bacteria at the Lower Size Limit of Life by Dan Krotz, Berkeley Lab (+Newswise), 27 Feb 2015. (See TEM image at right; scale bar = 100 nanometers.)
 Maria Lluch-Senar et al., "Defining a minimal cell: essentiality of small ORFs and ncRNAs in a genome-reduced bacterium" [Open Access abstract], doi:10.15252/msb.20145558, n 780 v 11, Mol Syst Biol, 21 Jan 2015. "We found that the minimal essential genome is comprised of 33% (269,410 bp) of the M. pneumoniae genome. Our data highlight an unexpected hidden layer of smORFs with essential functions, as well as non-coding regions, thus changing the focus when aiming to define the minimal essential genome." Maria Lluch-Senar et al., "Defining a minimal cell: essentiality of small ORFs and ncRNAs in a genome-reduced bacterium" [Open Access abstract], doi:10.15252/msb.20145558, n 780 v 11, Mol Syst Biol, 21 Jan 2015. "We found that the minimal essential genome is comprised of 33% (269,410 bp) of the M. pneumoniae genome. Our data highlight an unexpected hidden layer of smORFs with essential functions, as well as non-coding regions, thus changing the focus when aiming to define the minimal essential genome."
 10 Feb 2015: Life is a DNA software-driven system. 10 Feb 2015: Life is a DNA software-driven system.
 14 Jan 2015: Bacteria are like smartphones. 14 Jan 2015: Bacteria are like smartphones.
 Julie A Theriot, "Why are bacteria different from eukaryotes?" [html], doi:10.1186/1741-7007-11-119, n 119 v 11, BMC Biology, 13 Dec 2013. "If you go down the list of all the things that are special about eukaryotic cells, you can ascribe virtually all of them to functions of the cytoskeleton." Julie A Theriot, "Why are bacteria different from eukaryotes?" [html], doi:10.1186/1741-7007-11-119, n 119 v 11, BMC Biology, 13 Dec 2013. "If you go down the list of all the things that are special about eukaryotic cells, you can ascribe virtually all of them to functions of the cytoskeleton."
 What Is Life? A 21st Century Perspective by J. Craig Venter on the 70th Anniversary of Schroedinger's Lecture at Trinity College, 12 Jul 2012. "Life is code." What Is Life? A 21st Century Perspective by J. Craig Venter on the 70th Anniversary of Schroedinger's Lecture at Trinity College, 12 Jul 2012. "Life is code."
 13 Sep 2013: ...self-preservation... is the default existential condition of living things.... Pamela Lyon. 13 Sep 2013: ...self-preservation... is the default existential condition of living things.... Pamela Lyon.
 18 Apr 2013: Earth was seeded by panspermia. (Points to an article that discusses the complexity of prokaryotes.) 18 Apr 2013: Earth was seeded by panspermia. (Points to an article that discusses the complexity of prokaryotes.)
 10 May 2012: The definition of life and speculations about its origin(s)... by Gerald Joyce of the Scripps Research Institute. 10 May 2012: The definition of life and speculations about its origin(s)... by Gerald Joyce of the Scripps Research Institute.
 Edward N. Trifonov, "Opinion: What Is Life?" [html], The Scientist, 16 Feb 2012. "One could easily collect from the literature more than 100 different definitions, none satisfactory enough to be broadly accepted." Edward N. Trifonov, "Opinion: What Is Life?" [html], The Scientist, 16 Feb 2012. "One could easily collect from the literature more than 100 different definitions, none satisfactory enough to be broadly accepted."
 Michael Yarus, Chapter 8 from "Life from an RNA world: The ancestor within" [publisher's promo], Harvard University Press, Cambridge, MA, April, 2010. "...We could potentially store just over three hours of reasonably faithful music in the space afforded by a human genome." Michael Yarus, Chapter 8 from "Life from an RNA world: The ancestor within" [publisher's promo], Harvard University Press, Cambridge, MA, April, 2010. "...We could potentially store just over three hours of reasonably faithful music in the space afforded by a human genome."
 Sky Shields sends a link to a video with an extrapolation on the question, What Is Life?, 24 Mar 2011. Sky Shields sends a link to a video with an extrapolation on the question, What Is Life?, 24 Mar 2011.
 22 Feb 2011: We'll create panspermia.... — J. Craig Venter at a discussion of the definition of life. 22 Feb 2011: We'll create panspermia.... — J. Craig Venter at a discussion of the definition of life.
 Special series on the definition of life, p1001-1042 v10, Astrobiology, Dec 2010; and commentary: What is life? New answers to an age-old question in astrobiology, Physorg.com, 13 Jan 2011. Special series on the definition of life, p1001-1042 v10, Astrobiology, Dec 2010; and commentary: What is life? New answers to an age-old question in astrobiology, Physorg.com, 13 Jan 2011.
 Steven A. Benner, "Defining Life" [html], doi:10.1089/ast.2010.0524, Astrobiology, 16 Dec 2010. Steven A. Benner, "Defining Life" [html], doi:10.1089/ast.2010.0524, Astrobiology, 16 Dec 2010.
 Nick Lane and William Martin, "The energetics of genome complexity" [abstract], doi:10.1038/nature09486, p929-934 v467, Nature, 21 Oct 2010. "...Mitochondrial genes enabled a roughly 200,000-fold rise in genome size compared with bacteria." Nick Lane and William Martin, "The energetics of genome complexity" [abstract], doi:10.1038/nature09486, p929-934 v467, Nature, 21 Oct 2010. "...Mitochondrial genes enabled a roughly 200,000-fold rise in genome size compared with bacteria."
 Lowly bacteria are turning out to be much more complex than previously thought, Newswise, 12 Jul 2010. Lowly bacteria are turning out to be much more complex than previously thought, Newswise, 12 Jul 2010.
 Erika Check Hayden, "...Life is complicated" [html], doi:10.1038/464664a, Nature, 1 Apr 2010. Erika Check Hayden, "...Life is complicated" [html], doi:10.1038/464664a, Nature, 1 Apr 2010.
 5 Mar 2010: Genes in the microbes in our guts are 150 times more numerous than human genes. 5 Mar 2010: Genes in the microbes in our guts are 150 times more numerous than human genes.
 28 Nov 2009: There is no such a thing as a 'simple' bacterium — Howard Ochman and Rahul Raghavan. 28 Nov 2009: There is no such a thing as a 'simple' bacterium — Howard Ochman and Rahul Raghavan.
 28 Apr 2009: Life is nothing but an electron looking for a place to rest. 28 Apr 2009: Life is nothing but an electron looking for a place to rest.
 The Meaning of Life, by Carl Zimmer, SeedMagazine.com, 5 Sep 2007. "There is no one definition that we agree upon." The Meaning of Life, by Carl Zimmer, SeedMagazine.com, 5 Sep 2007. "There is no one definition that we agree upon."
 Smallest genome clocks in at 182 genes, doi:10.1038/news061009-10, News@Nature.com, 12 Oct 2006. Smallest genome clocks in at 182 genes, doi:10.1038/news061009-10, News@Nature.com, 12 Oct 2006.
 9 Feb 2006: A cell requires 430 genes, according to a new study from the J. Craig Venter Institute. 9 Feb 2006: A cell requires 430 genes, according to a new study from the J. Craig Venter Institute.
 20 Dec 2005: Life as We Do Not Know It, by Peter Ward [book review]. 20 Dec 2005: Life as We Do Not Know It, by Peter Ward [book review].
 17 Nov 2005: Hubert Yockey's definition of life (point #3) in email from his daughter, Cynthia Yockey. 17 Nov 2005: Hubert Yockey's definition of life (point #3) in email from his daughter, Cynthia Yockey.
 Victor Zykov et al., "Robotics: Self-reproducing machines" [abstract], p 163-164 v 435, Nature, 12 May 20005. Victor Zykov et al., "Robotics: Self-reproducing machines" [abstract], p 163-164 v 435, Nature, 12 May 20005.
 The Origin of Life on Earth re: Neil deGrasse Tyson and the definition of life, Astrobiology Magazine, online 29 Nov 2004. The Origin of Life on Earth re: Neil deGrasse Tyson and the definition of life, Astrobiology Magazine, online 29 Nov 2004.
 28 Sep 2004: Nova explains the origin of life. 28 Sep 2004: Nova explains the origin of life.
 K.C. Cole, "Seeking Life as We Know It" [text], Los Angeles Times, 5 Mar 2004. K.C. Cole, "Seeking Life as We Know It" [text], Los Angeles Times, 5 Mar 2004.
 Nick Campbell, "I can't live without you" [link], re: Bacillus subtilis's 271 essential genes, p 405 v 4, Nature Reviews Genetics, June 2003. Nick Campbell, "I can't live without you" [link], re: Bacillus subtilis's 271 essential genes, p 405 v 4, Nature Reviews Genetics, June 2003.
 Life's Working Definition: Does It Work? Astrobiology Magazine, 14 Apr 2003. Life's Working Definition: Does It Work? Astrobiology Magazine, 14 Apr 2003.
 Minimalist Life, by Stephen Hart, Astrobiology Magazine, 18 Dec 2002. Minimalist Life, by Stephen Hart, Astrobiology Magazine, 18 Dec 2002.
 2002, December 13: The Economist explains bioinformatics. 2002, December 13: The Economist explains bioinformatics.
 Natalie Angier, "Defining the Undefinable: The Living Cell" [text], The New York Times, 18 Dec 2001. Natalie Angier, "Defining the Undefinable: The Living Cell" [text], The New York Times, 18 Dec 2001.
 Living dead by Duncan Graham-Rowe, New Scientist Online News, 16 May 2001. "Ants and infertile humans are not alive, but parasitic DNA is.... A new, universal definition of life." Living dead by Duncan Graham-Rowe, New Scientist Online News, 16 May 2001. "Ants and infertile humans are not alive, but parasitic DNA is.... A new, universal definition of life."
 It's Not Just DNA Anymore: Prion Proteins and Hereditary Information, by Rabiya S. Tuma, issue 81, HMS Beagle, 23 June 2000. "The data now show that some proteins can carry hereditary information as well." It's Not Just DNA Anymore: Prion Proteins and Hereditary Information, by Rabiya S. Tuma, issue 81, HMS Beagle, 23 June 2000. "The data now show that some proteins can carry hereditary information as well."
 The Meaning of "Life", by Steven M. Potter, 11 March 1986. [6-page philosophical and historical analysis of definition of life written for Prof. Stanley Miller's Biochemical Evolution course.] The Meaning of "Life", by Steven M. Potter, 11 March 1986. [6-page philosophical and historical analysis of definition of life written for Prof. Stanley Miller's Biochemical Evolution course.]
 TIGR's Minimal Genome Project: How Many Genes Are Necessary to Sustain Life?, by Vicki Brower, HMS Beagle, 12 May 2000. TIGR's Minimal Genome Project: How Many Genes Are Necessary to Sustain Life?, by Vicki Brower, HMS Beagle, 12 May 2000.
 1999, December 10: Mimimal life requires about 300 genes. 1999, December 10: Mimimal life requires about 300 genes.
 Life: What Exactly Is It?, an Internet discussion among Stanley L. Miller, Antonio Lazcano, Mark Bedau, Thomas Ray and Claire Fraser, moderated by Miranda Ran in HMS Beagle, 25 June 1999. Life: What Exactly Is It?, an Internet discussion among Stanley L. Miller, Antonio Lazcano, Mark Bedau, Thomas Ray and Claire Fraser, moderated by Miranda Ran in HMS Beagle, 25 June 1999.
 1999, May 28: Dr. Baruch Blumberg challenges the scientific community "to look for life without any specifications." 1999, May 28: Dr. Baruch Blumberg challenges the scientific community "to look for life without any specifications."
 1999, April 12: A computer model of a cell requires a minimum of 127 genes. 1999, April 12: A computer model of a cell requires a minimum of 127 genes.
References
1. J.B.S. Haldane, What Is Life?, New York: Boni and Gaer, 1947. p 53.
2. Lynn Margulis, Symbiotic Planet: A New Look at Evolution, Basic Books, 1998. p 69. Also see: Chaper 1, "What Is Life?" in Origins of Sex by Lynn Margulis and Dorion Sagan, Yale Unversity Press, 1986.
3. Martin Olomucki, The Chemistry of Life. McGraw-Hill, Inc., 1993. p 25.
4. Harold J. Morowitz, The Beginnings of Cellular Life: Metabolism Recapitulates Biogenesis, Yale University Press, 1992. p 12.
4.2. Sydney Brenner, "Life's code script," Nature, 22 Feb 2012.
4.5. Daniel C. Dennett, Darwin's Dangerous Idea: Evolution and the Meanings of Life, New York, Simon & Schuster, Inc., 1995.
5. J. William Schopf, "The Oldest Fossils and What They Mean" p 29-63, Major Events in the History of Life, J. William Schopf, ed., Jones and Bartlett Publishers, 1992.
6. C. R. Woese, O. Kandler and M. L. Wheelis, "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya" [abstract], p4576-4579 v87, PNAS, June 1990.
7. Michael W. Gray, "The third form of life," p 299 v 383 Nature, 26 September 1996.
8. Hans-Peter Klenk, Lixin Zhou and J. Craig Venter, "Understanding life on this planet in the age of genomics," in Instruments, Methods, and Missions for the Investigation of Extraterrestrial Microorganisms, Richard B. Hoover, ed., Proceedings of SPIE Vol. 3111, p 306-317 (1997).
9. Carl R. Woese, and Norman R. Pace, "Probing RNA Structure, Function, and History by Comparative Analysis," p 91-117. The RNA World, R.F. Gesteland and J.F. Atkins, eds., Cold Spring Harbor Laboratory Press, 1993.
10. Leslie E. Orgel, The Origins of Life: Molecules and Natural Selection, John Wiley and Sons, Inc., 1973. p 92-93.
11. Denis Noble interviewed by Suzan Mazur, Huffpost, 2014.
![]()
|



